Bioinformatics - protocols B
To locate the required tools, look for a blue B
in the list of links or use the shortcuts
provided.
Basic sequence manipulation
You have just sequenced the following Arabidopsis cDNA fragment.
Note the format of the sequence: a header line (>mysequence) followed
by the rest of the sequence, which means FASTA format.
>cDNA_B_At
CCGCGGTGGCGGCCGCTCTAGAACTAGTGGATCCCCCGGGCTGCAGGATC
CCGGTGACCCCGGCAAAGCTTGCTTAATCCGAAGACGTTTCTGTTTCATC
TTCTTAAATCCGGGCCAACNGCGTTTACGAGACTAAACGCGTTTCTCTTT
AGGGCTTAATTATTATCCAGAGATGGCTCATCATAGCAAATGTTTACAAA
CGTTGGATTTAGCTTGTAAAGAGCTGAGATCTCGTGGCTTGTTTGTGAAG
CTTTTGGAGGCAATACTTAAAGCTGGAAACAGAATGAACGCGGGTACCGC
GAGAGGAAACGCTCAAGCGTTTAATCTAACCGCGCTTTTGAAGCTTTCGG
ATGTTAAAAGCGTTGATGGGAAGACTTCTTTGCTTAACTTTGTAGTGGAG
GAAGTTGTTAGATCGGAAGGAAAACGTTGTGTTATGAATAGAAGAAGCCA
TAGCTTAACACGAAGCGGTAGTAGTAACTACAATGGTGGTAATAGTAGTC
TTCAGGTTATGTCGAAAGAAGAGCAAGAGAAAGAGTACTTGAAGCTTGGT
TTACCAGTTGTTGGTGGATTGAGCTCTGAGTTTTCAAACGTGAAGAAAGC
TGCTTGTGTGGACTATGAAACGGTTGTTGCAACTTGTTCTGCTCTTGCGG
TTAGAGCGAAAGATGCGAAAACGGTGATTGGAGAATGTGAAGATGGAGAA
GGAGGGAGGTTTGTGAAAACGATGATGACGTTTCNTGATTCGGTAGAGGA
AGAGGTGAAAATAGCGAAAGGTGAAGAGAGGAAAGTGATCCCTGA
-
Look at the coding capacity of your cDNA using either the BCM
sequence utilities 6 Frame Translation function, or the
ExPASy
Translation Tool . Make sure that you are NOT using the default "Verbose"
option on ExPASy (or just try it once so that you know why avoiding it
:-). (You might want to look also at other translation tools such
as The Protein Machine).
-
Select the longest reading frame in the sense orientation and keep it as
a FASTA file. (NOTE: you might not necessarily get a reasonable ORF - assume
that this was a raw first run sequence, containing possible frameshifts).
Sequence similarity searches and domain structure
analysis
-
Use the protein sequence from the previous excercise as a query for a standard
protein BLAST search of the non-redundant
NCBI database. Uncheck the "Do CD Search option", otherwise keep default
parameters. If you were unable to perform the
search for server-dependent reasons, go to the SOS page.
You
should be able to produce a working hypothesis about the possible function
of your cDNA already at this stage!
-
From the BLAST results page, retrieve the protein sequence corresponding
to the best hit (by clicking the link to the left from the gene description-
see Figure, which is accidentaly from a nucleotide BLAST but the file structure
is the same). If the BLAST server is down, go
to the SOS page.
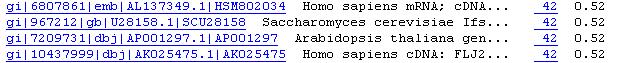
-
Retrieve the corresponding protein sequence in FASTA format (or,
in the worst case, from the SOS page).
Run a protein BLAST search as above using the retrieved protein sequence
as query and all default parameters including the CD-search.
-
On the 1st screen (before "Format Results") you will see a summary of conserved
domains found in your sequence by CD-search.
KEEP
THIS PAGE OPEN FOR FUTURE USE. If you lose it, you have to redo the search.
If the server is down, go to the SOS
page.
-
On the BLAST results page, look up which kind of things the search has
picked by looking at some of the alignments (links to the right from sequence
description). Note also the number of entries corresponding to all hits
with E-values better (i.e. lower) than 10-4 (denoted as e-04)
and repeat the BLAST search, disabling this time the low complexity filter
(you can uncheck CD-search as well for increased performance). Again, look
at the results and note the number of hits with E lower than 10-4.
Why is filtering the query a good idea? (Both
BLAST results can be found on the SOS page).
-
Perform a search for transmembrane or signal peptide sequences in your
hypothetical protein using
SignalP
and/or TMHMM.
Construction and interpretation of a protein sequence
alignment
Go back to the window with the CD-search results and
click one the conserved domain(s) found.
-
On the resulting page (backup on SOS)
chose "add query to the alignment", keeping all other parameters default,
and examine the output (could be found also on
the SOS page).Note how the identical sequence
picked from the database was (mis)aligned.
-
Repeat the previous task using the "most diverse set" and "top of CD search"
options.
Below you find the sequences that have been used to produce the first ("most
similar 10") alignment from above. Save then into a text file with extension
*.aa and create an alignment using MACAW.
-
Open MACAW on your computer (should
be OK over the browser). Go File... New Project...,
select Sequence type= protein and
Significance=number.
Chose the PAM120 scoring matrix and import sequence (Sequence...
Import...) from the file you just created. Save the Macaw file
on your computer before proceeding further in order to decrease the likelihood
of crash.
-
Examine the performance of Gibbs sampler vs. Segment pair overlap methods
and construct an alignment of the protein sequences.
-
(Optional): repeat the last task using a different
scoring matrix.
>Bprotein
SGGETSKQVKLKPLHWDKVNPDSDHSMVWDKIDRGSFSFDGDLMEALFGYVAVGKKSPEQ
GDEKNPKSTQIFILDPRKSQNTAIVLKSLGMTREELVESLIEGNDFVPDTLERLARIAPT
KEEQSAILEFDGDTAKLADAETFLFHLLKSVPTAFTRLNAFLFRANYYPEMAHHSKCLQT
LDLACKELRSRGLFVKLLEAILKAGNRMNAGTARGNAQAFNLTALLKLSDVKSVDGKTSL
LNFVVEEVVRSEGKRCVMNRRSHSLTRSGSSNYNGGNSSLQVMSKEEQEKEYLKLGLPVV
GGLSSEFSNVKKAACVDYETVVATCSALAVRAKDAKTVIGECEDGEGGRFVKTMMTFLDS
VEEEVKIAKGEERKVMELVKRTTDYYQAGAVTKGKNPLHLFVIVRDFLAMVDKVCLDIMR
NMQRRK
>gi|6691125 Nicotiana tabacum NFH2
EKSEEILKPKLKTLHWDKVRASSDCEMVWDQLKSSSFKLNEEMIETLFVVKNPTLNTSAT
AKHFVVSSMSQENRVLDPKKSQNIAILLRVLNGTTEEICEAFLEGNAENIGTELLEILLK
MAPSKEEERKLKEYKDDSPFKLGPAEKFLKAVLDIPFAFKRIDAMLYISNFDYEVDYLGN
SFETLEAACEELRSSRMFLKLLEAVLKTGNRMNVGTNRGDAHAFKLDTLLKLVDVKGADG
KTTLLHFVVQEIIKSEGARLSGGNQNHQQSTTNDDAKCKKLGLQVVSNISSELINVKKSA
AMDSEVLHNDVLKLSKGIQNIAEVVRSIEAVGLEESSIKRFSESMNRFMKVAEEKILRLQ
AQETLAMSLVKEITEYVHGDSAREEAHPFRIFMVVKDFLMILDCVCKEVGTINERTI
>gi|6041849 Arabidopsis BAC F21O3
EGTTDRPKPKLKPLPWDKVRPSSRRTNTWDRLPYNSSNANSKQRSLSCDLPMLNQESKVL
DPRKSQNVAVLLTTLKLTTNDVCQALRDGHYDALGVELLESLARVAPSEEEEKKLISYSD
DSVIKLAPSERFLKELLNVPFVFKRVDALLSVASFDSKVKHLKRSFSVIQAACEALRNSR
MLLRLVGATLEAGMKSGNAHDFKLEALLGLVDIKSSDGRTSILDSVVQKITESEGIKGLQ
VVRNLSSVLNDAKKSAELDYGVVRMNVSKLYEEVQKISEVLRLCEETGHSEEHQWWKFRE
SVTRFLETAAEEIKKIEREEGSTLFAVKKITEYFHVDPAKEEAQLLKVFVIVRDFLKILE
GVCKKMEVTSSLA
>gi|6225268 DIAPHANOUS PROTEIN HOMOLOG 1
PKKLYKPEVQLRRPNWSKLVAEDLSQDCFWTKVKEDRFENNELFAKLTLTFSAQTKTKKD
QEGGEEKKSVQKKKVKELKVLDSKTAQNLSIFLGSFRMPYQEIKNVILEVNEAVLTESMI
QNLIKQMPEPEQLKMLSELKDEYDDLAESEQFGVVMGTVPRLRPRLNAILFKLQFSEQVE
NIKPEIVSVTAACEELRKSESFSNLLEITLLVGNYMNAGSRNAGAFGFNISFLCKLRDTK
STDQKMTLLHFLAELCENDYPDVLKFPDELAHVEKASRVSAENLQKNLDQMKKQISDVER
DVQNFPAATDEKDKFVEKMTSFVKDAQEQYNKLRMMHSNMETLYKELGEYFLFDPKKLSV
EEFFMDLHNFRNMFLQAVKENQKRRKTEEKMRRAKLAKEKAEKERLEKQQKREQLIDMNA
EGDETGVMDSLLEALQSGAAFRR
>gi|544344 FORMIN 4 (LIMB DEFORMITY PROTEIN).
RKPAIEPSCPMKPLYWTRIQINDKSQDAAPTLWDSLEEPHIRDTSEFEYLFSKDTTQQKK
KPLSEAYEKKNKVKKIIKLLDGKRSQTVGILISSLHLEMKDIQQAIFTVDDSVVDLETLA
ALYENRAQEDELTKIRKYYETSKEEDLKLLDKPEQFLHELAQIPNFAERAQCIIFRAVFS
EGITSLHRKVEIVTRASKGLLHMKSVKDILALILAFGNYMNGGNRTRGQADGYSLEILPK
LKDVKSRDNGMNLVDYVVKYYLRYYDQEAGTDKSVFPLPEPQDFFLASQVKFEDLLKDLR
KLKRQLEASEQQMKLVCKESPREYLQPFKDKLEEFFKKAKKEHKMEESHLENAQKSFETT
VGYFGMKPKTGEKEVTPSYVFMVWFEFCSDFKTIWKRESKNISKER
>gi|2281090 unknown protein [Arabidopsis thaliana]
EKKVETMKPKLKTLHWDKVRASSSRVMVWDQIKSNSFQVNEEMIETLFKVNDPTSRTRDG
VVQSVSQENRFLDPRKSHNIAILLRALNVTADEVCEALIEGNSDTLGPELLECLLKMAPT
KEEEDKLKELKDDDDGSPSKIGPAEKFLKALLNIPFAFKRIDAMLYIVKFESEIEYLNRS
FDTLEAATGELKNTRMFLKLLEAVLKTGNRMNIGTNRGDAHAFKLDTLLKLVDIKGADGK
TTLLHFVVQEIIKFEGARVPFTPSQSHIGDNMAEQSAFQDDLELKKLGLQVVSGLSSQLI
NVKKAAAMDSNSLINETAEIARGIAKVKEVITELKQETGVERFLESMNSFLNKGEKEITE
LQSHGDNVMKMVKEVTEYFHGNSETHPFRIFAVVRDFLTILDQVCKEVGRVNERTV
>gi|1061334 Drosophila melanogaster cappuccino
RKSAVNPPKPMRPLYWTRIVTSAPPAPRPPSVANSTDSTENSGSSPDEPPAANGADAPPT
APPATKEIWTEIEETPLDNIDEFTELFSRQAIAPVSKPKELKVKRAKSIKVLDPERSRNV
GIIWRSLHVPSSEIEHAIYHIDTSVVSLEALQHMSNIQATEDELQRIKEAAGGDIPLDHP
EQFLLDISLISMASERISCIVFQAEFEESVTLLFRKLETVSQLSQQLIESEDLKLVFSII
LTLGNYMNGGNRQRGQADGFNLDILGKLKDVKSKESHTTLLHFIVRTYIAQRRKEGVHPL
EIRLPIPEPADVERAAQMDFEEVQQQIFDLNKKFLGCKRTTAKVLAASRPEIMEPFKSKM
EEFVEGADKSMAKLHQSLDECRDLFLETMRFYHFSPKACTLTLAQCTPDQFFEYWTNFTN
DFKDIWKKEITSLLNEL
>gi|5080823 Hypothetical protein [Arabidopsis thaliana]
GKTEDPTQPKLKPLHWDKMNPDASRSMVWHKIDGGSFNFDGDLMEALFGYVARKPSESNS
VPQNQTVSNSVPHNQTYILDPRKSQNKAIVLKSLGMTKEEIIDLLTEGHDAESDTLEKLA
GIAPTPEEQTEIIDFDGEPMTLAYADSLLFHILKAVPSAFNRFNVMLFKINYGSEVAQQK
GSLLTLESACNELRARGLFMKLLEAILKAGNRMNAGTARGNAQAFNLTALRKLSDVKSVD
AKTTLLHFVVEEVVRSEGKRAAMNKNMMSSDNGSGENADMSREEQEIEFIKMGLPIIGGL
SSEFTNVKKAAGIDYDSFVATTLALGTRVKETKRLLDQSKGKEDGCLTKLRSFFESAEEE
LKVITEEQLRIMELVKKTTNYYQAGALKERNLFQLFVIIRDFLGMVDNACSEIARNQRKQ
Q
Gene building: searching for coding sequences in
chromosomal DNA
The following DNA sequence corresponds to the sense strand of the locus
in the A. thaliana Chromosome 1 from which your initial cDNA has been derived:
>locusB
cccctataaaaagtattaaaaaggactgatacaataatgtatataaatat
cctaaaagatcttaattttgtaaatttattgttgtatattctaaacccgc
aatattagaatgatgatttagtaaacaagaaagacaaaataaataattaa
ttttagctagaaaagatgaaataaacactcatgatttaagccatacaaat
cgaagccccttgggttcagcatttctcaccaagtaaataccatcacctct
ggaaacccatttacgtacttgaccacatcttttattagcggctcctctgt
atgctctccatatgttatacacactatgatgccttaagatttattcacga
cgatttaatcagatacgcttatggattgccaaagatgatgccatctactt
agagaaaaacaatggaaagcgagaacgcatgtataattggaataaaaatt
aatatggttttcatatatctaaaaaattggacatttgaagccttaataaa
ttatactatgtaaaaatacttgtttatgaatgtaaattataataaattac
gatttaattagggaaatattgactatatatttcacccaaatattgaatgt
aaattttattttccaatacttttgcacatttaagaaattttcggatgtat
ttcctaaagaatattaccttttttgttttttaaaccatgcctttttgttt
tacacgttcataaatgcatgttccatacgcattaccataatttaatttga
acttaattttctctaggaatggtgatgatccactaccactatcattgatt
tcattccatattcctttgaccgactgaaattacgttggaaatagtatatt
ttgatgaataatttatttactcggaaaaaagaggtcaagttattaatagt
aagtacatatacattatcaattaagaattcaattgagttttaaggaaaat
cctattaatttgtttggtattcggtatttgttagttctaaggaattgaat
ttcccgattatacatcattataacgttctcaagttccaaacttgcaaccc
acattttgtcgatattctcaaatgtgaattcattcaatttcccatagaaa
acataaatttgcacttaaagttaacaattgaaatcgtatctaaatgggaa
tgtttttggcttttagtgttagacttccaaagcgtcaaaaatatttctag
aaagagcacaaaaaataagcaacgccactacttttggacaaagtcaacga
taacacacatcaaccgcaccagctccataaaagtccatctcacgaaaacg
attctagtcaaactacctaaaacacccttatatttacatacaacccaatc
ccactaacaagggtattttcgtcaatcacaaaatttatcaccgacccggg
aagaagaagaagaacagatcaactaatttctgctttcaactccacattaa
accaaaacctccaaaaagaatcatttatttaaattatcttcccgttttaa
gttcctgagatttttgggaattgtaaatttgaagaaaattaaacaaagac
gtgttttcatttttttttttgtttcctttattgatctctctctatctctc
taaatgagctaaatcgttaatggctgccatgtttaatcatccatggccta
atttaaccctaatttacttcttcttcatcgtcgttttaccattccaatca
ctttctcaatttgattctcctcaaaatatcgaaactttcttccccatctc
ttcactctcccctgttccaccaccgcttcttccaccttcgtcaaacccat
ctccgccgtcgaataattcatcatcttcggataaaaaaacaatcaccaaa
gctgtccttataacagcagcaagtactttacttgtagctggagttttctt
cttctgcctccaaagatgtatcatcgcacggagacggagagacagagttg
gaccagtcagagtcgaaaacactttacctccgtatcctcctcctccgatg
acgtcggcggcggtgactacgactactttggctagagaaggattcacgag
gtttggtggtgtgaaaggtttgattcttgatgagaatggtcttgatgtgt
tgtattggagaaagctacagagtcagagagaaagaagtgggagtttcagg
aaacagatcgtcaccggagaagaagaagacgagaaagaagttatttatta
caagaacaagaagaaaacagagcccgttacagagattcctcttcttagag
gaagatcatctacttctcacagtgttatccataacgaagatcatcagccg
ccaccgcaggtgaaacagagtgaaccaacaccaccaccgccaccaccgtc
aattgcggtgaaacagagtgcaccaacgccatcgccacctcctccgatta
agaagggttcttcaccatcgccaccgccacctccaccggtgaaaaaggtt
ggagctttatcatcatcagcttcgaaaccaccacctgcgccggttagagg
agcaagtggaggagagacttcgaaacaagtaaagttgaagcctttacatt
gggataaagtaaaccctgattccgatcattcaatggtttgggacaaaatc
gatcgtggatcattcaggtatatatttatttcgaaagttagggcttttgc
ttcaatcaattgaaaaaaccctaatttgtttttgtttcttctcagtttcg
atggcgatttaatggaagctctgtttggatacgttgccgtggggaagaaa
tcaccagaacaaggcgatgagaaaaaccctaaatcaacgcaaatattcat
acttgatccgagaaagtctcaaaacacagcgattgtgctcaaatcattag
gtatgacacgtgaagagcttgttgaatcactcatagaaggaaacgatttc
gtgccagacactcttgagaggttagctagaatagctccaacgaaagaaga
acaatcagccattcttgaattcgacggtgacacggcaaagcttgctgatg
cggagacgtttctgtttcatcttcttaaatccgtgccaaccgcgtttacg
agactaaacgcgtttctctttagggctaattattatccagagatggctca
tcatagcaaatgtttacaaacgttggatttagcttgtaaagagctgagat
ctcgtggcttgtttgtgaagcttttggaggcaatacttaaagctggaaac
agaatgaacgcgggtaccgcgagaggaaacgctcaagcgtttaatctaac
cgcgcttttgaagctttcggatgttaaaagcgttgatgggaagacttctt
tgcttaactttgtagtggaggaagttgttagatcggaaggaaaacgttgt
gttatgaatagaagaagccatagcttaacacgaagcggtagtagtaacta
caatggtggtaatagtagtcttcaggttatgtcgaaagaagagcaagaga
aagagtacttgaagcttggtttaccagttgttggtggattgagctctgag
ttttcaaacgtgaagaaagctgcttgtgtggactatgaaacggttgttgc
aacttgttctgctcttgcggttagagcgaaagatgcgaaaacggtgattg
gagaatgtgaagatggagaaggagggaggtttgtgaaaacgatgatgacg
tttcttgattcggtagaggaagaggtgaaaatagcgaaaggtgaagagag
gaaagtgatggagcttgtgaaacgtacaacggattattatcaagcaggag
ctgttacaaaggggaagaatccacttcatttgtttgttatcgttagagat
tttcttgccatggttgataaagtttgcttagatattatgagaaatatgca
gaggaggaaggttggtagtccgatatcgccttcttcgcagcggaatgcgg
tgaaattcccggttttgcctccgaatttcatgtcggacagagcttggagt
gattctggtgggtcggattctgatatgtgagagtcaagatttgttatatg
taaatactaaatagtagaagcattttgggtattgattagcattgaaagat
gttgaattgtttatagatttatcagtccaaagcattggacttgagtataa
tttgttccttgtataaataaacaattttgctttaagacctttccatgttt
atgaacatgtcttctttaacttcacatagaccttttgtttacgtaagaac
taataatactaaattgtttgataattctaaatgtgaaagtgaaccactat
atagtgtgaacttggctttattgaattctttttaaaaaaatttctccaga
gctttagatgtaggagttaatattttcacctaacatagcctcttttttat
gtttctctatcaactaacactaaatttgtggatgaagactaaattaacat
aagtttatctattaactaacaacctaccagtttgatgcttgtaaatatga
aacttcaacgttataaagactatatggtgtgaactttttatccatcttta
ttgacttttaaaattttcttaatttgagtaaacaaaagcagaagcttttt
aaaggatgcaggagttgatttttgtatatgaacaaaacatatacttctcc
cttagacgaatttggagctatcattcttggtttcaaactttttaataatt
tgagctttaaagcaaaatggcaactttatattgatcactagtccacaaca
ctttctctgccttttcctcaatagcaacgcgtagtcaagaagaagaacgt
gtttaacatggaccaatcttgattaagataatagtatgatcaaatgctta
tataaacacactaaaaaggaatcaaatttaaccattccacaaatcaccaa
caaaatttaatgaatcatgtctctgcttctaaagatgttattattttcct
tattcttcttctatatggcttcaatttctcaatgctcagacccaaccggt
ggacagtttagcttcaacggttacttgtacaccgatggagttgcggatct
aaacccggacggtttgttcaaactcataacttcaaagaca
-
Try to predict the exon-intron structure using the NetGene2
server (chose Arabidopsis, and take care to load only sequence, not the
FASTA header, into the bottom box). (If server
down, the results can be found on the SOS page.)
-
(Optional): try the same using the
Web Gene Gene Builder interface. Take into account the results
of the previous excercise (the simplest way is using the best protein match,
provided for your convenience below, as key protein). Chose Direct strand,
Gene Model, Sequence error report, Use EST mapping and Complete gene model;
switch off the Protein homology search. Don`t forget to select organism
= Arabidopsis, and the plant scoring matrix! (NOTE: there are currently
problems at the server, but look at least at the interface to get an idea
what it should do).
-
Look up results of other prediction methods - GenScan
and GeneFinder (use links or contact group A)
and compare the results.
Key protein sequence:
>gi|6691125|gb|AAF24497.1|AF213696_1 FH protein NFH2 [Nicotiana
tabacum]
MVFPFFFFLLFLFCSTHCISFAAVSAHNRRVLHESFFPIDSPPPSQPPIPAPPAPPTPYPFQPSTPDNNN
PFFPTYRSPPPPPPPPSPSSLVSFPANISDINLPNTSKSKHVSSKLIITAITCVLAAIIVLSIAICLHAK
KRRRHFNDPKTQRSDNSNRLNHGSSKNDGNTNNSIPKLQQPSQTSSEFLYLGTIVNSHGGINSGSNPDTA
PSSRKMASPELRPLPPLNGRNLSQNYRNTRNDDDFYSTEESVGYIESSFGAGSLSRRGFAAVEVNKFVGS
SLSGSDSSSSSGSGSPNRSVSLSISPPVSVSPKRESCSRPKSPELIAVVTPPPPQRPPPPPPPFVHGPQV
KVTANESPVLISPMEKNDQNVENHSIEKNEEKSEEILKPKLKTLHWDKVRASSDCEMVWDQLKSSSFKLN
EEMIETLFVVKNPTLNTSATAKHFVVSSMSQENRVLDPKKSQNIAILLRVLNGTTEEICEAFLEGNAENI
GTELLEILLKMAPSKEEERKLKEYKDDSPFKLGPAEKFLKAVLDIPFAFKRIDAMLYISNFDYEVDYLGN
SFETLEAACEELRSSRMFLKLLEAVLKTGNRMNVGTNRGDAHAFKLDTLLKLVDVKGADGKTTLLHFVVQ
EIIKSEGARLSGGNQNHQQSTTNDDAKCKKLGLQVVSNISSELINVKKSAAMDSEVLHNDVLKLSKGIQN
IAEVVRSIEAVGLEESSIKRFSESMNRFMKVAEEKILRLQAQETLAMSLVKEITEYVHGDSAREEAHPFR
IFMVVKDFLMILDCVCKEVGTINERTIVSSAQKFPVPVNPNLQPVISGFRAKRLHSSSDEESSSP